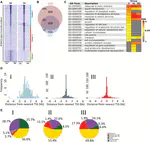
long non-coding RNA (lncRNA)
Over the past decades, advances in RNA sequencing methods (RNA-Seq) have revealed that a large proportion of eukaryotic genomes are transcribed outside the coding regions. These thousands of new transcripts have been named non-coding RNAs (ncRNA). Among them, long ncRNAs (lncRNAs) are RNA longer than 200nt with a low protein coding capacity. They are generally polyadenylated, lowly expressed and in a fairly localised manner (ie in few cells) or specific in response to environmental stresses.
LncRNAs have long been regarded as transcriptional noise due to their low protein coding potential, their poor conservation and their low level of expression. However, an increasing number of studies have shown that lncRNAs can regulate gene expression at different levels and play an important role in a variety of biological processes. LncRNAs have been shown to regulate the expression of contiguous genes (cis action) or act at a distance (trans action) via interactions with ribonucleoproteins or other protein complexes, modulate splicing of mRNA transcripts and/or hijack proteins or RNA molecules. Therefore they seems to be involved in all steps of gene expression regulation.
According to their locations with respect to the other genes, lncRNAs are classified into different categories:
- intergenic lncRNAs (lincRNAs), when they are located in between two genes
- intronic lncRNAs, when they are located in introns of coding genes
- sense lncRNAs, when they are the result of specific RNA isoform of a coding genes
- natural antisense transcripts (NATs), when they are located on the other strand of the DNA of a gene.
The major challenges in lncRNAs investigation are to understand their mechanism of actions and how are they evolving.